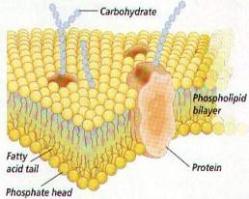
Our research activities fall within 3 main areas, which all relate to the study of the kinetics and thermodynamics of protein conformational changes, namely membrane protein folding, protein-detergent interactions and protein fibrillation. These areas are linked by a keen interest in understanding the mechanistic and thermodynamic behaviour of proteins in different circumstances by quantifying the strength of internal side-chain interactions as well as contacts with solvent molecules, whether it be detergents, denaturants, stabilizing salts and osmolytes or lipids. Ultimately we hope this will lead to a greater manipulative ability vis-a-vis processes of both basic, pharmaceutical and industrial relevance. The general approach is to use available spectroscopic techniques (fluorescence, CD, stopped-flow, FTIR, NMR and dynamic and static light scattering) to generate data which can be analyzed in a quantitative manner to develop models and mechanisms for conformational changes at the molecular level.