iSMS implements two methods for calculating fluorescence intensities of individual molecules:
The main disadvantages of using Gaussian PSF fitting is the inherent requirement on a good fit of each frame, which is often unreliable at low signal-to-noise ratios. Additionally, the fit depends on starting parameters and constraints. This means that Gaussian PSF does not work equally well on all molecules. Secondly, Gaussian PSF fitting is somewhat more slow compared to aperture photometry. Thirdly, the molecular PSF is not necessarily Gaussian shaped.
The main advantage of Gaussian PSF fitting is a lower fluctuation of background at high intensities and the ability to follow the Gaussian parameters in time, which is the technique exploited in TFM.
You can compare the performance of the different methods as described below.
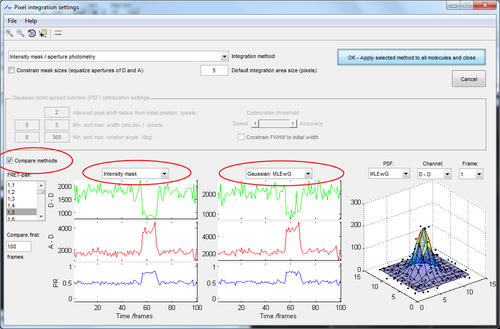
If the integration aperture (mask) of a molecule does not fit perfectly on to the molecule you can interactively define the integration mask of the molecule in the FRET-pair window.